



1.250 Å
X-ray
2007-04-02
| Name: | Phenylalanine 2-monooxygenase precursor |
|---|---|
| ID: | PAO_PSESP |
| AC: | Q5W9R9 |
| Organism: | Pseudomonas sp |
| Reign: | Bacteria |
| TaxID: | 306 |
| EC Number: | 1.13.12.9 |
| Chain Name: | Percentage of Residues within binding site |
|---|---|
| B | 100 % |
| B-Factor: | 8.371 |
|---|---|
| Number of residues: | 69 |
| Including | |
| Standard Amino Acids: | 63 |
| Non Standard Amino Acids: | 0 |
| Water Molecules: | 6 |
| Cofactors: | |
| Metals: | |
| Ligandability | Volume (Å3) |
|---|---|
| 0.924 | 938.250 |
| % Hydrophobic | % Polar |
|---|---|
| 39.57 | 60.43 |
| According to VolSite | |
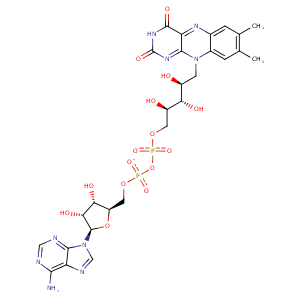
| HET Code: | FAD |
|---|---|
| Formula: | C27H31N9O15P2 |
| Molecular weight: | 783.534 g/mol |
| DrugBank ID: | DB03147 |
| Buried Surface Area: | 80.97 % |
| Polar Surface area: | 381.7 Å2 |
| Number of | |
|---|---|
| H-Bond Acceptors: | 22 |
| H-Bond Donors: | 7 |
| Rings: | 6 |
| Aromatic rings: | 3 |
| Anionic atoms: | 2 |
| Cationic atoms: | 0 |
| Rule of Five Violation: | 3 |
| Rotatable Bonds: | 13 |
| X | Y | Z |
|---|---|---|
| 71.755 | 34.4788 | 91.549 |

 Image generated by PoseView
Image generated by PoseView



Represent the protein/ligand binding mode, centered on the ligand
Dashed lines represents hydrogen bonds and metal interactions
Green residue labels for amino acids with hydrophobic contacts (green lines) to the ligand

| Ligand | Protein | Interaction | |||
|---|---|---|---|---|---|
| Atom | Atom | Residue | Distance (Å) | Angle (°) | Type |
| C4' | CB | ALA- 66 | 4.26 | 0 | Hydrophobic |
| O1P | N | GLY- 67 | 2.86 | 152.3 | H-Bond (Protein Donor) |
| O3B | OE2 | GLU- 94 | 2.67 | 169.56 | H-Bond (Ligand Donor) |
| O3B | OE1 | GLU- 94 | 3.22 | 124.09 | H-Bond (Ligand Donor) |
| O2B | OE1 | GLU- 94 | 2.61 | 164.2 | H-Bond (Ligand Donor) |
| N3A | N | ALA- 95 | 3.22 | 152.03 | H-Bond (Protein Donor) |
| O1A | N | ARG- 119 | 2.84 | 165.84 | H-Bond (Protein Donor) |
| O2A | NH2 | ARG- 119 | 3.36 | 136.4 | H-Bond (Protein Donor) |
| O2A | NE | ARG- 119 | 2.9 | 165.53 | H-Bond (Protein Donor) |
| O3P | NH2 | ARG- 119 | 3.01 | 132.47 | H-Bond (Protein Donor) |
| O2A | CZ | ARG- 119 | 3.59 | 0 | Ionic (Protein Cationic) |
| C8M | CD | ARG- 119 | 3.94 | 0 | Hydrophobic |
| C9 | CB | ARG- 119 | 4.29 | 0 | Hydrophobic |
| C3' | CB | ARG- 119 | 4.47 | 0 | Hydrophobic |
| C9A | CB | ALA- 141 | 4.18 | 0 | Hydrophobic |
| C2' | CB | ALA- 141 | 4.13 | 0 | Hydrophobic |
| O4 | N | MET- 142 | 2.91 | 165.96 | H-Bond (Protein Donor) |
| N3 | O | ARG- 143 | 2.83 | 149.78 | H-Bond (Ligand Donor) |
| O4 | N | ARG- 143 | 2.77 | 154.92 | H-Bond (Protein Donor) |
| N6A | O | VAL- 374 | 2.93 | 157.42 | H-Bond (Ligand Donor) |
| N1A | N | VAL- 374 | 3 | 156.35 | H-Bond (Protein Donor) |
| C1B | CG1 | VAL- 410 | 4.18 | 0 | Hydrophobic |
| C7M | CB | SER- 476 | 4.27 | 0 | Hydrophobic |
| C7M | CE2 | TYR- 536 | 4.02 | 0 | Hydrophobic |
| C7M | CE2 | TRP- 608 | 4.46 | 0 | Hydrophobic |
| C8M | CE2 | TRP- 608 | 3.62 | 0 | Hydrophobic |
| C7M | CE2 | PHE- 617 | 4.43 | 0 | Hydrophobic |
| C8M | CD2 | PHE- 617 | 4.49 | 0 | Hydrophobic |
| O2P | OG | SER- 651 | 2.72 | 159.59 | H-Bond (Protein Donor) |
| O3' | OD2 | ASP- 652 | 2.82 | 176.62 | H-Bond (Ligand Donor) |
| O2P | N | ASP- 652 | 3.08 | 154.2 | H-Bond (Protein Donor) |
| N1 | N | LEU- 661 | 3.37 | 131.36 | H-Bond (Protein Donor) |
| O2 | N | LEU- 661 | 2.91 | 164.96 | H-Bond (Protein Donor) |
| C4' | CD2 | LEU- 661 | 3.8 | 0 | Hydrophobic |
| C2' | CD2 | LEU- 661 | 3.99 | 0 | Hydrophobic |
| C5' | CB | ALA- 664 | 4.08 | 0 | Hydrophobic |
| O1A | O | HOH- 1906 | 2.63 | 156.05 | H-Bond (Protein Donor) |
| O1P | O | HOH- 1917 | 2.71 | 168.32 | H-Bond (Protein Donor) |
| O2 | O | HOH- 1945 | 2.87 | 179.96 | H-Bond (Protein Donor) |
| O3B | O | HOH- 1964 | 2.77 | 147.67 | H-Bond (Protein Donor) |