



2.190 Å
X-ray
2011-07-11
| Name: | Polyprotein |
|---|---|
| ID: | A8DG50_9HEPC |
| AC: | A8DG50 |
| Organism: | Hepatitis C virus subtype 1a |
| Reign: | Viruses |
| TaxID: | 31646 |
| EC Number: | / |
| Chain Name: | Percentage of Residues within binding site |
|---|---|
| B | 74 % |
| C | 26 % |
| B-Factor: | 32.809 |
|---|---|
| Number of residues: | 44 |
| Including | |
| Standard Amino Acids: | 43 |
| Non Standard Amino Acids: | 0 |
| Water Molecules: | 1 |
| Cofactors: | |
| Metals: | |
| Ligandability | Volume (Å3) |
|---|---|
| 1.231 | 1312.875 |
| % Hydrophobic | % Polar |
|---|---|
| 40.10 | 59.90 |
| According to VolSite | |
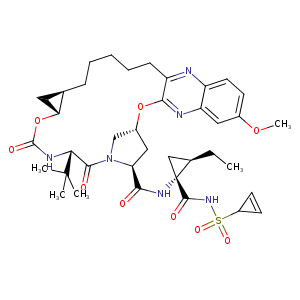
| HET Code: | SUE |
|---|---|
| Formula: | C38H50N6O9S |
| Molecular weight: | 766.903 g/mol |
| DrugBank ID: | DB11575 |
| Buried Surface Area: | 61.27 % |
| Polar Surface area: | 203.59 Å2 |
| Number of | |
|---|---|
| H-Bond Acceptors: | 11 |
| H-Bond Donors: | 3 |
| Rings: | 7 |
| Aromatic rings: | 2 |
| Anionic atoms: | 0 |
| Cationic atoms: | 0 |
| Rule of Five Violation: | 2 |
| Rotatable Bonds: | 8 |
| X | Y | Z |
|---|---|---|
| -16.4016 | 9.5132 | 1.99357 |
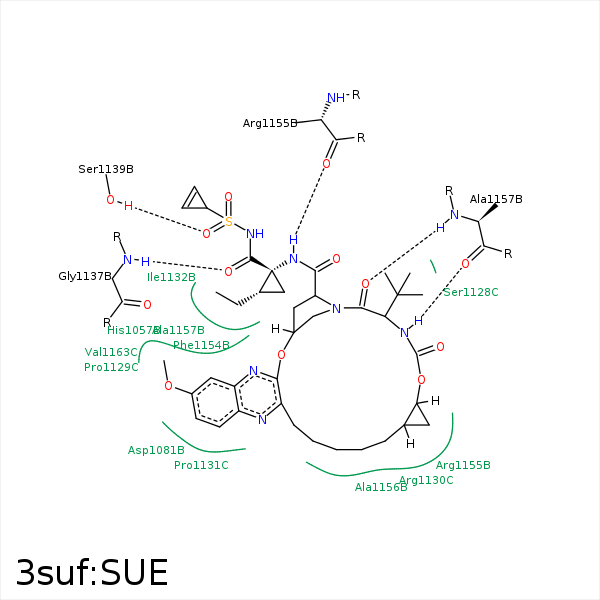
 Image generated by PoseView
Image generated by PoseView



Represent the protein/ligand binding mode, centered on the ligand
Dashed lines represents hydrogen bonds and metal interactions
Green residue labels for amino acids with hydrophobic contacts (green lines) to the ligand

| Ligand | Protein | Interaction | |||
|---|---|---|---|---|---|
| Atom | Atom | Residue | Distance (Å) | Angle (°) | Type |
| CBA | CE1 | TYR- 1056 | 3.92 | 0 | Hydrophobic |
| NAV | NE2 | HIS- 1057 | 3.24 | 134.66 | H-Bond (Ligand Donor) |
| CBA | CB | HIS- 1057 | 4.31 | 0 | Hydrophobic |
| CBX | CB | HIS- 1057 | 4.48 | 0 | Hydrophobic |
| CAE | CB | HIS- 1057 | 3.68 | 0 | Hydrophobic |
| CAY | CG2 | VAL- 1078 | 4.07 | 0 | Hydrophobic |
| CAD | CB | ASP- 1081 | 3.63 | 0 | Hydrophobic |
| CAR | CB | SER- 1128 | 4.42 | 0 | Hydrophobic |
| CG1 | CB | SER- 1128 | 4.2 | 0 | Hydrophobic |
| CAE | CB | PRO- 1129 | 4 | 0 | Hydrophobic |
| CAD | CG | PRO- 1131 | 3.23 | 0 | Hydrophobic |
| CBQ | CG2 | ILE- 1132 | 3.42 | 0 | Hydrophobic |
| CG2 | CG2 | ILE- 1132 | 4.38 | 0 | Hydrophobic |
| CAU | CZ | TYR- 1134 | 3.65 | 0 | Hydrophobic |
| CAS | CB | LEU- 1135 | 3.91 | 0 | Hydrophobic |
| OBK | N | GLY- 1137 | 3.25 | 124.79 | H-Bond (Protein Donor) |
| OBM | N | GLY- 1137 | 3.21 | 149.61 | H-Bond (Protein Donor) |
| CBN | CB | SER- 1139 | 3.91 | 0 | Hydrophobic |
| CBN | CZ | PHE- 1154 | 3.33 | 0 | Hydrophobic |
| CBI | CG | ARG- 1155 | 3.96 | 0 | Hydrophobic |
| NBP | O | ARG- 1155 | 2.64 | 141.22 | H-Bond (Ligand Donor) |
| CAG | CB | ALA- 1156 | 4.05 | 0 | Hydrophobic |
| CAX | CB | ALA- 1156 | 3.4 | 0 | Hydrophobic |
| CAX | CB | ALA- 1156 | 3.4 | 0 | Hydrophobic |
| N | O | ALA- 1157 | 2.91 | 156.87 | H-Bond (Ligand Donor) |
| O | N | ALA- 1157 | 2.97 | 168.42 | H-Bond (Protein Donor) |
| CG2 | CB | ALA- 1157 | 4.25 | 0 | Hydrophobic |
| CBQ | CB | ALA- 1157 | 3.83 | 0 | Hydrophobic |
| CAF | CG1 | VAL- 1158 | 4.04 | 0 | Hydrophobic |
| CAY | CG2 | VAL- 1163 | 4.2 | 0 | Hydrophobic |
| CBA | CG1 | VAL- 1163 | 3.89 | 0 | Hydrophobic |
| CBJ | CG1 | VAL- 1163 | 3.9 | 0 | Hydrophobic |