



2.200 Å
X-ray
2006-04-05
| Name: | 1,5-anhydro-D-fructose reductase |
|---|---|
| ID: | AFR_ENSAD |
| AC: | Q2I8V6 |
| Organism: | Ensifer adhaerens |
| Reign: | Bacteria |
| TaxID: | 106592 |
| EC Number: | / |
| Chain Name: | Percentage of Residues within binding site |
|---|---|
| A | 2 % |
| B | 98 % |
| B-Factor: | 28.185 |
|---|---|
| Number of residues: | 40 |
| Including | |
| Standard Amino Acids: | 40 |
| Non Standard Amino Acids: | 0 |
| Water Molecules: | 0 |
| Cofactors: | |
| Metals: | |
| Ligandability | Volume (Å3) |
|---|---|
| 0.399 | 634.500 |
| % Hydrophobic | % Polar |
|---|---|
| 45.74 | 54.26 |
| According to VolSite | |
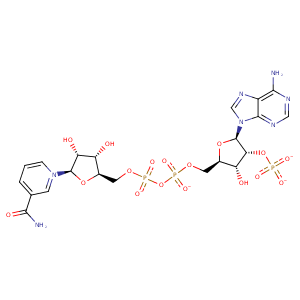
| HET Code: | NDP |
|---|---|
| Formula: | C21H26N7O17P3 |
| Molecular weight: | 741.389 g/mol |
| DrugBank ID: | DB02338 |
| Buried Surface Area: | 61.77 % |
| Polar Surface area: | 404.9 Å2 |
| Number of | |
|---|---|
| H-Bond Acceptors: | 22 |
| H-Bond Donors: | 5 |
| Rings: | 5 |
| Aromatic rings: | 2 |
| Anionic atoms: | 4 |
| Cationic atoms: | 0 |
| Rule of Five Violation: | 2 |
| Rotatable Bonds: | 13 |
| X | Y | Z |
|---|---|---|
| -4.16269 | 32.0236 | 72.9239 |
 Image generated by PoseView
Image generated by PoseView



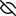
Represent the protein/ligand binding mode, centered on the ligand
Dashed lines represents hydrogen bonds and metal interactions
Green residue labels for amino acids with hydrophobic contacts (green lines) to the ligand

| Ligand | Protein | Interaction | |||
|---|---|---|---|---|---|
| Atom | Atom | Residue | Distance (Å) | Angle (°) | Type |
| O2X | N | ALA- 9 | 2.85 | 160.88 | H-Bond (Protein Donor) |
| C5D | CB | SER- 10 | 4.25 | 0 | Hydrophobic |
| O2A | N | THR- 11 | 2.88 | 142.55 | H-Bond (Protein Donor) |
| O2A | OG1 | THR- 11 | 2.68 | 162.37 | H-Bond (Protein Donor) |
| O1N | N | ILE- 12 | 2.97 | 167.01 | H-Bond (Protein Donor) |
| C5N | CB | ILE- 12 | 3.7 | 0 | Hydrophobic |
| C5D | CG2 | ILE- 12 | 3.95 | 0 | Hydrophobic |
| C4N | CD1 | ILE- 12 | 3.41 | 0 | Hydrophobic |
| O1X | OG | SER- 33 | 2.61 | 158.82 | H-Bond (Protein Donor) |
| O3X | OG1 | THR- 34 | 2.83 | 162.88 | H-Bond (Protein Donor) |
| O3X | N | THR- 34 | 3.19 | 155.38 | H-Bond (Protein Donor) |
| O1X | NH2 | ARG- 38 | 3.22 | 132.15 | H-Bond (Protein Donor) |
| O1X | NE | ARG- 38 | 2.86 | 145.8 | H-Bond (Protein Donor) |
| O1X | CZ | ARG- 38 | 3.45 | 0 | Ionic (Protein Cationic) |
| DuAr | CZ | ARG- 38 | 3.68 | 168.49 | Pi/Cation |
| C5B | CG2 | THR- 72 | 3.68 | 0 | Hydrophobic |
| O3D | OD1 | ASN- 73 | 2.78 | 149.38 | H-Bond (Ligand Donor) |
| C4D | CG | GLU- 93 | 4.11 | 0 | Hydrophobic |
| N7N | OE1 | GLU- 93 | 2.88 | 147.76 | H-Bond (Ligand Donor) |
| O2D | NZ | LYS- 94 | 3.08 | 153.44 | H-Bond (Protein Donor) |
| O2D | O | LYS- 94 | 2.64 | 142.42 | H-Bond (Ligand Donor) |
| C3N | CD | LYS- 94 | 4.17 | 0 | Hydrophobic |
| O1A | NE1 | TRP- 162 | 3.19 | 152.39 | H-Bond (Protein Donor) |
| C5D | CZ2 | TRP- 162 | 4.38 | 0 | Hydrophobic |
| C3D | CH2 | TRP- 162 | 3.81 | 0 | Hydrophobic |
| O2N | CZ | ARG- 163 | 3.57 | 0 | Ionic (Protein Cationic) |
| O2N | NH2 | ARG- 163 | 3.25 | 142.14 | H-Bond (Protein Donor) |
| O2N | NH1 | ARG- 163 | 3.02 | 156.27 | H-Bond (Protein Donor) |
| O7N | OH | TYR- 283 | 2.54 | 162.18 | H-Bond (Protein Donor) |